Single-cell metabolic profiling of human cytotoxic T cells
Hartmann, F.J., Mrdjen, D., McCaffrey, E.
et al.,
Nat Biotechnol (2020)
Key Insights
- Scientists at Stanford University, Vrije University Amsterdam, and the California Institute of Technology developed a method they call scMEP (short for single-cell metabolic regulome profiling) to quantify proteins associated with regulating the activity of metabolic pathways
- They paired scMEP with IONpath’s multiplexed ion beam imaging (MIBI™) technology to characterize immune cells
- They assessed and validated more than 100 antibodies for a broad range of metabolomic targets, selecting about 40 for this study and showing that the antibodies make it possible to infer immune cell identity in heterogeneous populations based on metabolomic regulome profiles
- For an average of 94% of the cells studied, metabolic features accurately predicted cell identity
- For each 400 μm × 400 μm image of tissue sample studied, scientists could interrogate 36 antibody dimensions and identify cell lineage, activation status, and more. A detailed analysis of the images revealed site-specific enzyme enrichment associated with a particular metabolic pathway, including respiratory, glycolysis, and amino acid pathways.
Abstract
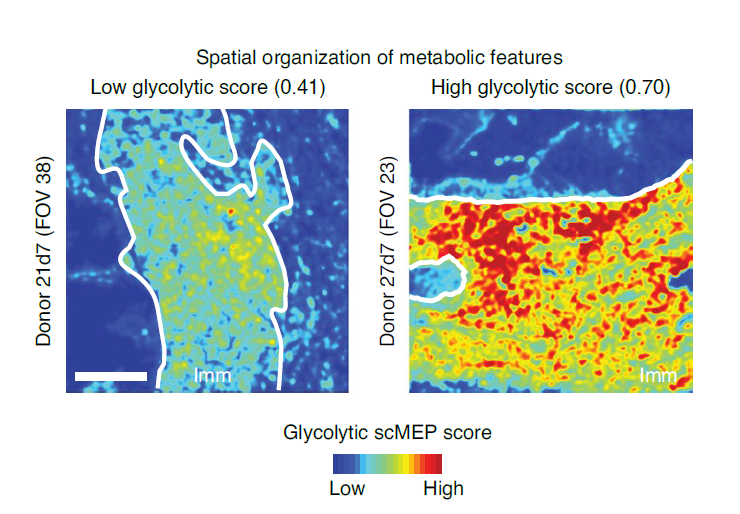
Cellular metabolism regulates immune cell activation, differentiation and effector functions, but current metabolic approaches lack single-cell resolution and simultaneous characterization of cellular phenotype. In this study, we developed an approach to characterize the metabolic regulome of single cells together with their phenotypic identity. The method, termed single-cell metabolic regulome profiling (scMEP), quantifies proteins that regulate metabolic pathway activity using high-dimensional antibody-based technologies. We employed mass cytometry (cytometry by time of flight, CyTOF) to benchmark scMEP against bulk metabolic assays by reconstructing the metabolic remodeling of in vitro-activated naive and memory CD8+ T cells. We applied the approach to clinical samples and identified tissue-restricted, metabolically repressed cytotoxic T cells in human colorectal carcinoma. Combining our method with multiplexed ion beam imaging by time of flight (MIBI-TOF), we uncovered the spatial organization of metabolic programs in human tissues, which indicated exclusion of metabolically repressed immune cells from the tumor–immune boundary. Overall, our approach enables robust approximation of metabolic and functional states in individual cells.
https://doi.org/10.1038/s41587-020-0651-8